1. INTRODUCTION
2. MATERIALS AND METHODS
2.1 Sample collection, DNA extraction, and metagenome sequencing
2.2 Analysis of assembly, gene prediction, functional annotations, and phylogeny
3. RESULTS AND DISCUSSION
3.1 Analysis of rhodolith metagenome sequences
3.2 Carbon and nitrogen cycling in rhodolith holobiont
4. CONCLUSION
1. INTRODUCTION
Rhodoliths are morphologically diverse and free-living non-geniculate coralline red algae (Rhodophyta), which are generally distributed in worldwide marine benthic habitats (Bosence 1983; Foster 2001; Fredericq et al. 2019; Jeong et al. 2019). Rhodolith beds are especially common from the low intertidal zone to the mesophotic zone (ca. ~150 m depth) along the continental shelf and near oceanic islands (Harris et al. 1996; Amado-Filho and Pereira-Filho 2012). An individual rhodolith is distinguished from crustose algae-coated pebble (McCoy and Kamenos 2015). The size and density of rhodoliths on the seafloor varies with water depth and latitude (Horta et al. 2016). Rhodoliths play an important ecological role by providing a microhabitat in a wide range of floral and faunal communities, e.g. kelp beds, corals, sponges, and herbivores, which are generally associated with biodiversity, abundance and food webs (Grall et al. 2006; Amado-Filho and Pereira-Filho 2012; Amado-Filho et al. 2017; Gabara 2020). For instance, a recent study showed numerous single-celled organisms including Prorocentrum lima (dinoflagellate) and Ochrosphaera verrucosa (haptophyte) are inhabiting within calcified cell lumens of rhodoliths using SEM, TEM and fluorescence micrographs (Krayesky-Self et al. 2017). Rhodolith beds also play crucial biogeochemical roles through photosynthesis and biomineralization of calcium carbonate (CaCO3) skeletons (Cabioch et al. 1999; Halfar et al. 2000; Williams et al. 2011; Kamenos and Law 2010; McCoy and Kamenos 2015). The depositions of calcified cell walls are constructed from dissolved inorganic carbon (DIC: aquatic CO2, HCO3-, and CO32-) and photosynthesis also responds to the elevated DIC in coralline algae (Gao et al. 1993; McCoy and Kamenos 2015). Carbonic anhydrases play crucial role in photosynthesis (Atkinson et al. 2016; Gee and Niyogi 2017; Razzak et al. 2019), and these enzymes are also involved in the calcifying mechanism of cnidarian coral skeletons (Bertucci et al. 2013). Rhodolith beds and crustose coralline algae are truly one of major contributors to carbonate deposition in the world’s oceans. These algae are hypothesized to be major contributors of the global ocean carbon biogeochemical cycling; however, biological interactions of rhodolith beds with global and local environmental changes are still unclear (Testa and Bosence 1999; Amado-Filho and Pereira-Filho 2012; Basso 2012; McCoy and Kamenos 2015; Horta et al. 2016; Teed et al. 2020).
Nitrogen metabolism is also involved in the carbon fixation mechanisms, and it is significantly important in algal growth (e.g., rhodolith beds) (Macler 1986; Huppe and Turpin 1994; Zhou et al. 2016; Schubert et al. 2019). However, usable nitrogen sources (e.g., ammonium, ammonia, nitrate) for eukaryotes are frequently acquired from microbial nitrogen fixers (N2-fixers); these diazotrophs include oxygenic/anoxygenic phototrophic, aerobic/anaerobic heterotrophic, bacteria, and archaea (Zhang et al. 2020). The epiphytic or symbiotic diazotrophs generate a nitrogen flux to rhodoliths and other macroalgal communities, and this nitrogen source leads to biological storage and cycling of both carbon and nitrogen within the marine trophic structure (Capone and Carpenter 1982; Phlips and Zeman 1990; Matson and Quenga 2005; Gabara 2020; Zehr and Capone 2020; Zhang et al. 2020). Rhodoliths and their microbial community consistently interact and form an ecological unit termed holobiont, which responds to environmental changes (Cavalcanti et al. 2018; van der Loos et al. 2019). The rhodolith holobionts include bacteria, archaea, virus, and diverse small eukaryotic organisms that were discovered by metagenomic approaches therefore rhodolith beds are ecologically important as hotspots of marine biodiversity, productivity, and resilience (Cavalcanti et al. 2013, 2014, 2018; Fredericq et al. 2019). However, biological interactions within the rhodolith holobiont are poorly studied.
To get more deeper understanding of rhodolith holobionts, we generated a metagenome sequencing data of a rhodolith using the whole-genome shotgun sequencing method, and we analyzed the species composition and its potential interactions. Our results reveal that the aerobic and anaerobic microbial community interact within the holobiont, playing an important role as the interactive channel for carbon and nitrogen cycling between biotic and abiotic environments around the rhodoliths.
2. MATERIALS AND METHODS
2.1 Sample collection, DNA extraction, and metagenome sequencing
The rhodolith samples were collected from Udo Island, Jeju, Korea (vicinity of 33°31'03.4"N 126°56'10.9"E; 2014.09.18; September 18, 2014), and placed in liquid nitrogen for several minutes. One of the frozen rhodolith samples was ground using a mortar and pestle, and total genomic DNA was extracted using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany). Whole-metagenome sequencing was conducted by Ion Torrent PGM platform with 400 bp-sized sequencing library kit (Thermo Fisher Scientific, San Francisco, California, USA). Raw reads (SRR12920572; https://www.ncbi.nlm.nih.gov/sra? linkname=bioproject_sra_all&from_uid=672827) and assembly (JADWZK000000000; https://www.ncbi.nlm.nih.gov/nuccore/ JADWZK000000000) of metagenome sequencing were uploaded to the NCBI SRA (Sequence Read Archive) database (BioProject PRJNA672827; https://www.ncbi.nlm.nih.gov/bioproject/PRJNA672827).
2.2 Analysis of assembly, gene prediction, functional annotations, and phylogeny
The sequenced metagenome raw reads were assembled by MEGAHIT assembler (Li et al. 2015). To classify the rhodolith sample, we analyzed 18S ribosomal DNA (rDNA) sequences using local BLASTn search (e-value cutoff = 1.e-10) with full length rDNA sequences of coralline algae (from Sporolithales, Rhodogorgonales, Hapalidiales, and Corallinales; Lee et al. 2018). The complete 18S rDNA sequences in the contig was analyzed by the web-based program RNAmmer 1.2 Server (Lagesen et al. 2007). Based on the assembly, protein sequences were analyzed by the Prodigal gene prediction software (Hyatt et al. 2010). Protein functions were analyzed by DIAMOND search (Buchfink et al. 2015) based on KEGG database (metabolic pathway analysis; http://www.genome.jp/tools/blast). The processes of assembly, gene prediction, and functional annotation were conducted by the SqueezeMeta pipeline (Tamames and Puente-Sánchez 2019). All BLAST and functional databases were downloaded using “download_databases.pl” script, which is involved in the SqueezeMeta pipeline (Tamames and Puente-Sánchez 2019). Taxonomic assignments of assembled contigs and analyzed genes in this study were defined by a representative taxa of BLAST/DIAMOND search results that analyzed by a fast LCA (lowest common ancestor; Luo et al. 2014) algorithm with nr database (Download: Aug. 2020) on the SqueezeMeta pipeline (Tamames and Puente-Sánchez 2019). The target gene was analyzed by BLASTp search (e-value cutoff = 1.e-05), and aligned with top 100 homologous sequences from the BLAST results using MAFFT v7.313 (default option: --auto; Yamada et al. 2016). Phylogenetic analysis was done using the maximum likelihood (ML) method with 1,000 bootstrap replications, and the best-fit evolutionary model was selected by default selection option (IQ-tree v1.6.12; Nguyen et al. 2015).
3. RESULTS AND DISCUSSION
3.1 Analysis of rhodolith metagenome sequences
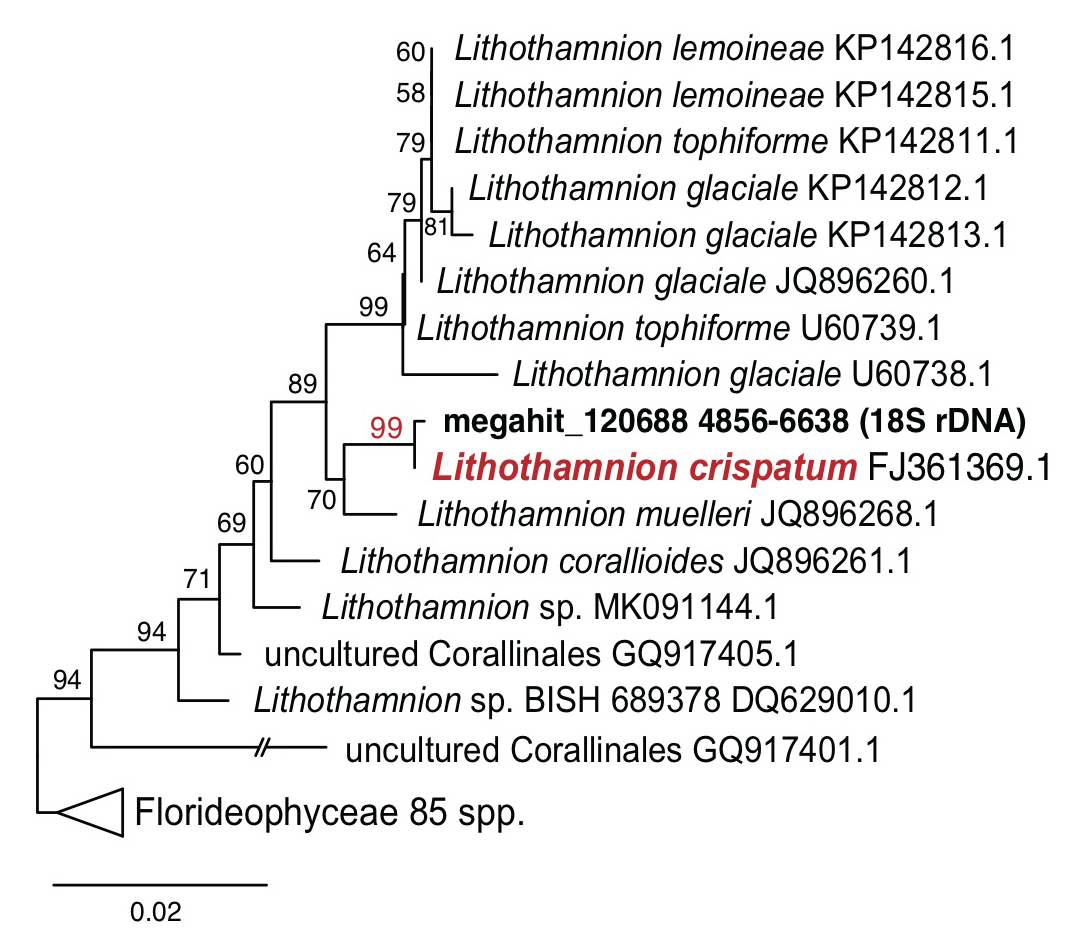
The generated rhodolith metagenome sequencing data (1.3 Gbp, the Ion Torrent PGM platform; Thermo Fisher Scientific, San Francisco, California) were assembled (210 Mbp; 376,038 contigs; N50=581 bp) by MEGAHIT assembler (Li et al. 2015). Based on the assembly, a total of 395,055 proteins was analyzed by the Prodigal gene prediction software (Hyatt et al. 2010). From the assembly, we found 18S ribosomal DNA (rDNA) sequences in contig ‘megahit_120688’, which is 1,783 bp in length, and this shows 99.9% sequence similarity and a strong monophyletic relationship with Lithothamnion crispatum (FJ361369.1; Fig. 1). A non-geniculate coralline alga, L. crispatum is a cosmopolitan rhodolith forming alga that is known to play an important role in biogenic calcium carbonate deposition (Basso et al. 2011; Amado-Filho et al. 2012; de Carvalho et al. 2017).
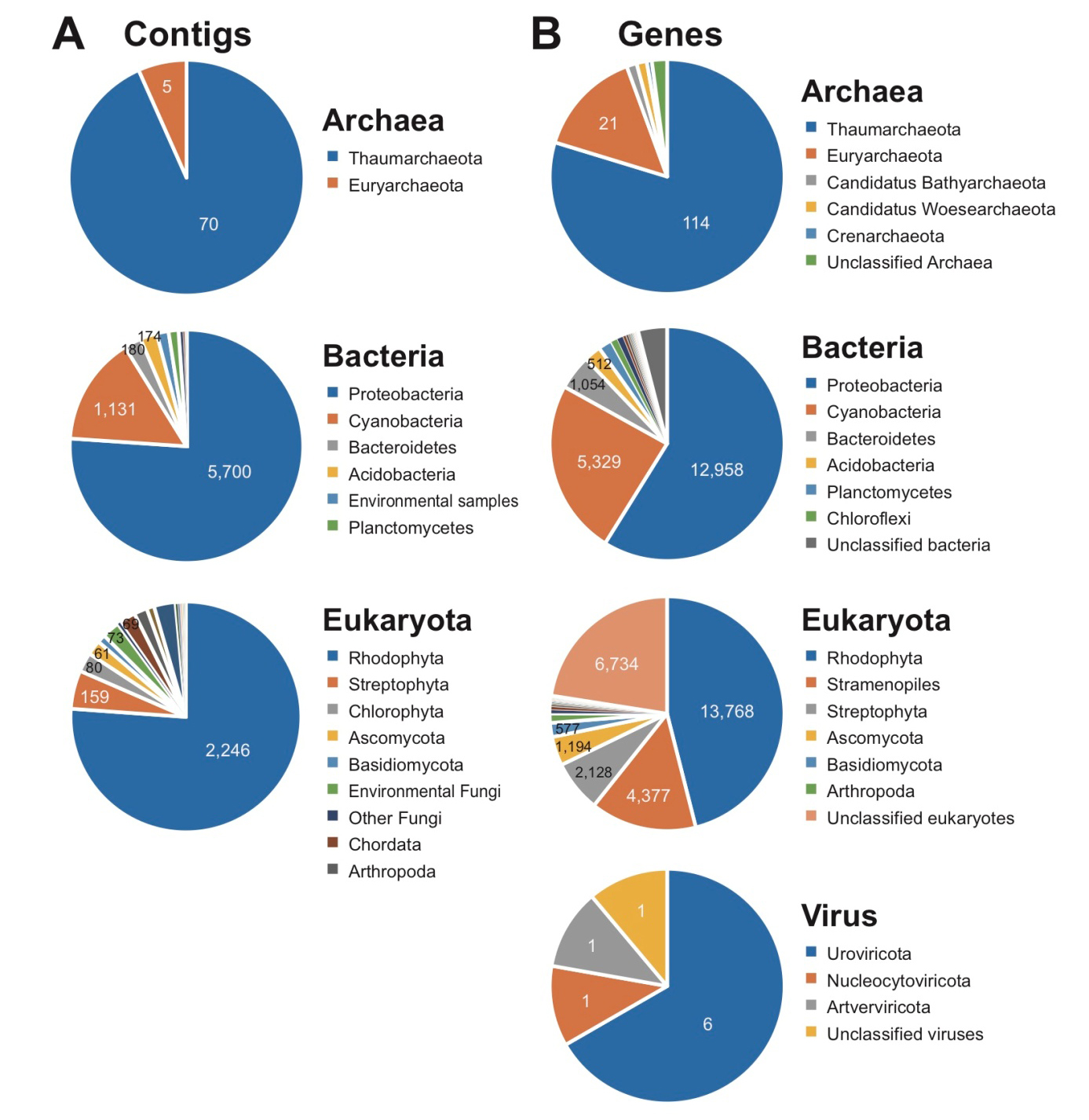
BLASTn analysis (e-value cutoff = 1.e-10) was done using 376,052 assembled contigs with nt database (Download: Oct. 2019), and a total of 10,541 contigs shows BLAST top hits to Archaea (75), Bacteria (7,493), and Eukaryota (2,950), Virus (1), and unclassified sequences (22), including Thaumarchaeota (70; Archaea), Proteobacteria (5,700; Bacteria), and Rhodophyta (2,246; Eukaryota), respectively (Fig. 2A). In eukaryotic contigs, fragmented plastid (1,127 contigs) and mitochondrial (702 contigs) genomes were found representing diverse taxonomic groups: identified plastid genome fragments from Rhodophyta (1,039), Chlorophyta (41), Stramenopiles (32), Streptophyta (11), Cryptophyta, (2), Euglenophyta (1), unclassified plastid (1) and mitochondrial genome fragments from Rhodophyta (661), Metazoa (16), Stramenopiles (9), Streptophyta (6), Fungi (6), Jakobida (2), Chlorophyta (1), Amoebozoa (1). Interestingly, BLAST top match sequences of fragmented 16 plastid contigs in our metagenome assembly belong to the chlorophyte algae Ostreobium sp. (KY819067, KY766991, KY766993, KY766995, and KY766996) and O. quekettii (LT593849); Ostreobium is the most abundant endolithic algal genus in marine environments, especially in cnidarian coral holobionts (Marcelino and Verbruggen 2016; Verbruggen et al. 2017; Massé et al. 2018). Ostreobium is also known to be a major species for biogenic dissolution of carbonates from coral reefs to the open ocean, which may have a significant impact on the calcium carbonate budget of coral reefs (Aline 2008; Grange et al. 2015). Our results suggest that Ostreobium may also contribute to the bioerosion of carbonates in rhodolith holobionts.
A total of 395,055 genes was analyzed in the metagenome assembly, and 55,399 genes were annotated based on taxonomic assignments of the analyzed genes (Supplementary Table 1; Fig. 2B). Although the metagenome data cannot represent quantification of species abundance, the taxonomic composition of the genes includes Archaea (143), Bacteria (22,016), and Eukaryota (29,896), Viruses (9), and unclassified taxa (3,335), and their major taxa are consistent with the results using contigs as follows: Thaumarchaeota (114; Archaea), Proteobacteria (12,958; Bacteria), and Rhodophyta (13,768; Eukaryota) (Fig. 2B). However, the results of taxonomic annotations using the genes show a more diverse taxonomic composition than those for the contigs (Fig. 2A and 2B). Among Rhodophyta genes, most of them (94.2%) were identified as the Florideophyceae that includes the subclass Corallinophycidae (i.e., coralline algae). The Bangiophyceae genes (2.3%) are present, indicating filamentous and foliose seaweeds (Blouin et al. 2011; Kucera and Saunders 2012; Koh and Kim 2018). Our rhodolith could include unicellular red algae (i.e., class Porphyridiophyceae and Cyanidiophyceae; genes < 2%, respectively among red algal genes). Because only several red algal genomes are available in the NCBI database, species-level compositional studies require additional red algal genomes. The Streptophyta genes (Supplementary Table 1) are regarded as seagrass genes but some of genes could be other algal genes because many genes in primary endosymbiosis groups show monophyletic relationships with other sister algal lineages (Chan et al. 2011). Other, diverse photosynthetic algal lineages are also present; brown algae (229 genes), diatoms (206 genes), green algae (51 genes), dinoflagellates (13 genes), haptophytes (8 genes), and apicomplexans (8 genes). This result corresponds well with natural habitats of Ecklonia cava and Undaria peterseniana (Hwang et al. 2010). Several dinoflagellates and haptophytes have a benthic stage (e.g., cysts) that is endolithically associated with rhodolith holobionts (Krayesky-Self et al. 2017). We found 13 genes of Scleractinia group (stony corals), which provide potential evidence for physical interactions between calcifying coralline algae and cnidarian corals. Cnidarian corals frequently contain symbionts (e.g., Symbiodinium) or parasites (e.g., apicomplexan-related lineage) (Janouškovec et al. 2012, 2013; González-Pech et al. 2019). Regardless of whether the apicomplexan genes in our metagenome data are directly associated with the rhodolith cells or the coral cells, it is evident that complex communities form the holobiont. Interestingly, oomycetes (water molds; non-photosynthetic Stramenopiles; 3,985 genes) shows relatively larger proportion in the rhodolith metagenome data. The oomycetes are indicated as fungi-like pathogens or parasites in plant hosts, which are physiologically related each other (Birch et al. 2006; Thines and Kamoun 2010; Richards et al. 2011; Pais et al. 2013). We suggest that rhodoliths also have a host-parasite interaction.
3.2 Carbon and nitrogen cycling in rhodolith holobiont
We focused on carbon and nitrogen cycle genes from the functional annotations of genes. A total of 25 carbonic anhydrases (CAs) was found in the rhodolith holobiont: 20 are regarded as bacterial, four as eukaryotic, and one with an unknown origin (Supplementary Table 2). The CA genes have an important role of capturing CO2, which is used in crucial cellular processes (e.g., photosynthesis) called carbon concentrating mechanism (Badger and Price 1994; Supuran and Capasso 2017; Wang et al. 2020). Diverse photosynthetic organisms in the rhodolith holobiont are involved in carbon cycling through the photosynthetic carbon fixation. Especially, coralline algae (i.e., rhodolith) may actively use the mechanism to deposit carbon sources not only through their plastid (photosynthesis) but also calcified cell walls (calcification) based on the dissolved inorganic carbon sources such as another calcifying organism, cnidarian corals (Gao et al. 1993; Bertucci et al. 2013; McCoy and Kamenos 2015; Wang et al. 2020).
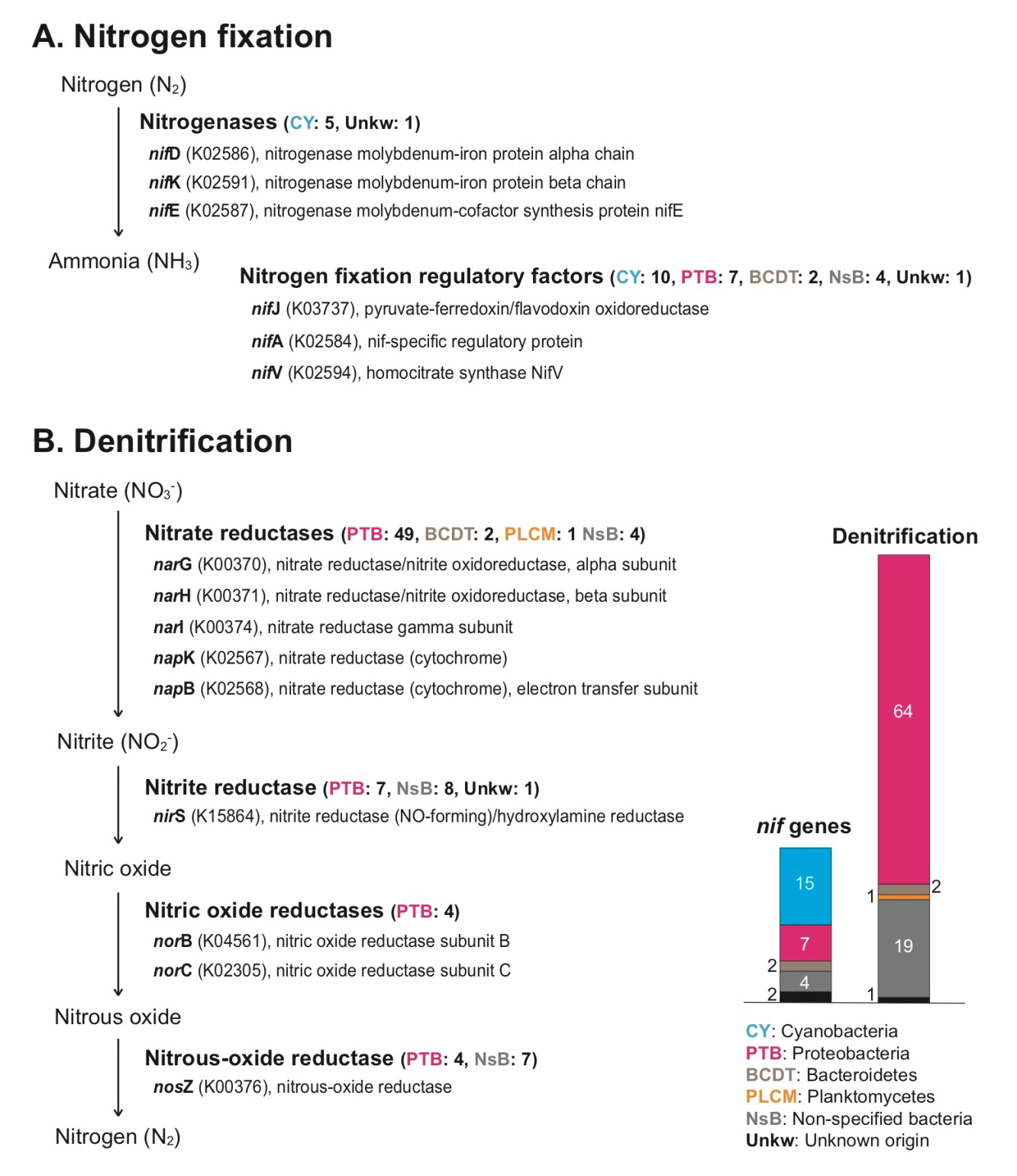
We found the following diverse genes involving nitrogen fixation (nif): nifD (K02586) and nifK (K02591) for nitrogenase molybdenum-iron protein function, nifE (K02587) for nitrogenase molybdenum-cofactor synthesis protein, nifJ (K03737) for pyruvate-ferredoxin/flavodoxin oxidoreductase, nifA (K02584) for nif-specific regulatory protein, nifV (K02594) for homocitrate synthase protein, and nif11 domain (pfam07862) containing protein (Supplementary Table 3). Among them, nifD, nifK, and nifE genes have core function (nitrogenase) during nitrogen fixation, directly converting nitrogen gas (N2) to ammonia (NH3) (Fani et al. 2000). Most of nitrogenase genes and their regulatory factors in the rhodolith holobiont were cyanobacterial genes (Fig. 3A, and Supplementary Table 3). In marine environments, biological nitrogen fixation of microbial communities is an important factor to supply usable nitrogen sources as key nutrients to algal cells through assimilation, and the nitrogen fixers are present in diverse algal holobionts (Cheng 2008; Barott et al. 2011; Sohm et al. 2011; de Oliveira et al. 2012; Egan et al. 2013). Several non-cyanobacterial (e.g., proteobacterial) nif regulatory factors were also present in the rhodolith holobiont (Fig. 3A, and Supplementary Table 3), and these are possibly indicated as non-cyanobacterial nitrogen fixers (Riemann et al. 2010; Farnelid et al. 2011; Sohm et al. 2011; Delmont et al. 2018). Based on these findings, we suggest that microbial nitrogen fixers in rhodolith populations contribute to nitrogen cycling. Interestingly, there are many different kinds of denitrification genes that include nitrate reductases (narG, narH, narI, napK, and napB), nitrite reductase (nirS), nitric oxide reductases (norB, and norC), and nitrous-oxide reductase (nosZ) (Fig. 3B, and Supplementary Table 4). Most denitrification genes are identified as proteobacterial genes (i.e., alphaproteobacterial, gammaproteobacterial, deltaproteobacterial, and unidentified proteobacterial genes, see Supplementary Table 4). Proteobacteria groups are abundant in diverse environments, and play an important ecological role in carbon, sulfur, and nitrogen cycles. Among the proteobacterial groups, deltaproteobacteria include diverse anaerobic families (Greene 2014; Kuever 2014a, 2014b). Several strict anaerobic deltaproteobacterial families including Desulfobacteraceae, Desulfobulbaceae, and Desulfuromonadaceae (strict but some tolerance to oxygen) are also present in the rhodolith holobiont, and they have nitrogen fixation and denitrification functions (Supplementary Table 3, and 4; Greene 2014; Kuever 2014a, 2014b). Indeed, it was reported that anaerobic sediments were preserved in rhodolith beds (Adey et al. 2015). We suggest the rhodolith holobiont generally has an anaerobic zone too, and their anaerobic bacterial community contributes the nitrogen cycling along with other aerobic/aerotolerant populations. The diffusion of oxygen in the particle microenvironments can be limited by the diffusive boundary layer more than nitrate, and anaerobic nitrogen metabolisms can occur in the central area of the microenvironment (Bianchi et al. 2018). Therefore, the re-cycled nitrogen may occur between the nitrogen fixation and denitrification organisms within the rhodolith holobiont (i.e., the inside of crustose algae-coated pebble including deposition of calcium carbonate). However, actual measurements of carbon and nitrogen flows in rhodolith are still required because our metagenomics approach can only reveal functional potential and cannot distinguish dormant microorganisms (Lennon and Jones 2011).
4. CONCLUSION
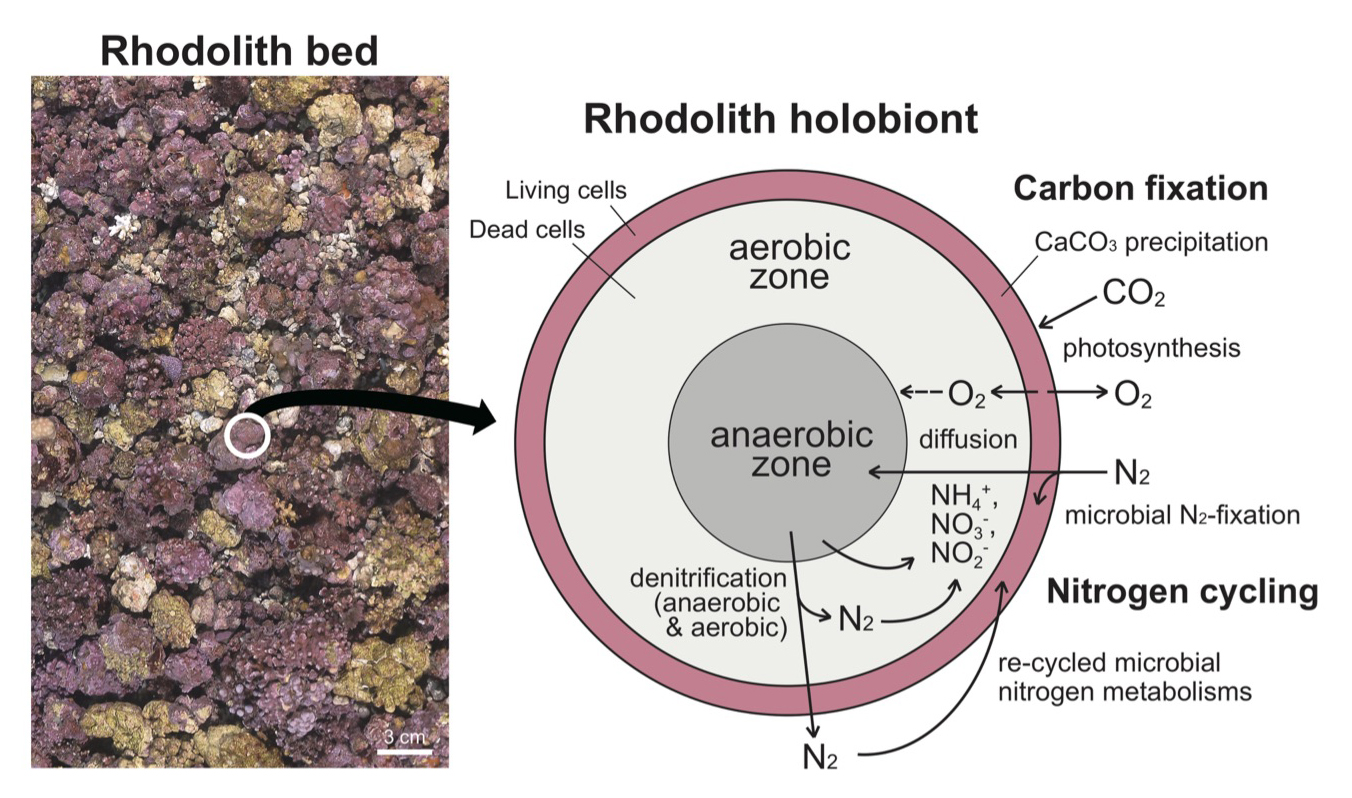
Rhodoliths (coralline algae) may support crucially for carbon cycling through calcium carbonate precipitations in their cell walls and photosynthesis, even though the cellular activities are only present in living cells of surface area (Fig. 4). Cyanobacterial communities also make photosynthetic activities in the photic surface of rhodoliths, and they conduct nitrogen fixation mechanisms (Supplementary Table 2 and 3), which could use in cell growth of rhodolith as well as other epiphytic algal community. The active interaction between cyanobacteria and rhodolith algae could accelerate carbon cycling. On the other hand, dead cells were filled inside of rhodolith (Fig. 4). These calcified dead cells within rhodolith serve habitats for diverse microbials communities as seedbanks and temporal reservoirs (Krayesky-Self et al. 2017; Fredericq et al. 2019). Because we found strict anaerobic microbial community (i.e., Desulfobacteraceae, Desulfobulbaceae) from our metagenome data as well as deltaproteobacterial species from rhodolith holobionts in a previous metagenome study (Cavalcanti et al. 2018), it is likely that the rhodolith holobiont comprises of aerobic and anaerobic zones as a functionally enclosed biosphere (Fig. 4). Interestingly, both aerobic and anaerobic denitrifications occur in rhodolith holobiont, and these re-cycled nitrogen sources may be used in nitrogen fixation again within the holobiont. The internal microbial interactions within the rhodolith holobiont may have advantages to maintain their biodiversity and respond directly to local environmental changes as self-sustaining reservoirs for marine carbon and nitrogen cycling. More metagenome data from rhodoliths could clarify the heterogeneity of microhabitats (e.g., surface and interior parts; Kim et al. 2021), and also reveal the major functions of the community structures (Amado-Filho and Pereira-Filho 2012; Amado-Filho et al. 2017; Cavalcanti et al. 2018).