1. Introduction
2. Materials and Methods
2.1 Sample Collection and Morphological Analysis
2.2 Molecular Analysis
2.3 Phylogenetic and Biogeographical Analysis
3. Results and Discussion
1. Introduction
The family Syngnathidae comprises approximately 330 species worldwide (Fricke et al. 2025). This group shows unique morphological and physiological characteristics, including male pregnancy, dermal plates enclosing the body, and an elongated snout. In particular, the location of the male brood pouch (on the trunk or tail) is considered a key morphological character that distinguishes two major phylogenetic lineages of the family (Hamilton et al. 2017). These characteristics are crucial for understanding evolution and phylogenetic relationships at the genus and species levels. Currently, 17 species in nine genera of syngnathid fishes have been recorded from Korea, and their identification is primarily based on morphological characters such as the number of body rings, fin-ray counts, head ridge structure, and color patterns (Lee and Kim 2020; Yoon et al. 2022).
Halicampus punctatus (Kamohara 1952), the starry pipefish, is known to occur mainly in continental shelf waters influenced by the Tsushima Current, particularly around Okinawa and Kyushu, Japan (Dawson 1985; Senou 2002; Sonoyama et al. 2020). This species is a tail-brooding pipefish (Urophori), characterized by a brood pouch situated on the tail (Breder and Rosen 1966). In the waters surrounding the Korean Peninsula, the sole record to date is that of larvae collected in the southwestern East Sea in December (Kim et al. 2006). However, because syngnathid larvae are planktonic and may be transported considerable distances by prevailing currents, the presence of larvae alone does not necessarily indicate the local occurrence or establishment of adult populations. In addition, larval morphology varies substantially across ontogenetic stages, and species-diagnostic characters are often not fully developed, making accurate morphological identification challenging (Ko et al. 2013; Okiyama 2014). Consequently, larval records alone are insufficient to confirm the establishment of this species in Korean waters, and adult specimens are required to definitively verify its distribution.
Meanwhile, molecular phylogeny for the genus Halicampus remains limited. A re-examination of COI sequences available in NCBI/GenBank (Clark et al. 2016) revealed significant interspecific genetic distances within the genus, suggesting potential species misidentifications of the genus in some records. These results suggested that the genus Halicampus required detailed molecular phylogenetic analyses and taxonomic treatment.
In this context, the present study re-examines the phylogenetic position of H. punctatus based on morphological identification and COI sequence analysis of five adult specimens collected for the first time from the eastern waters off Jeju Island. This study provides the first record of adult H. punctatus from Korean waters and offers baseline data for the taxonomic reassessment of the genus Halicampus.
2. Materials and Methods
2.1 Sample Collection and Morphological Analysis
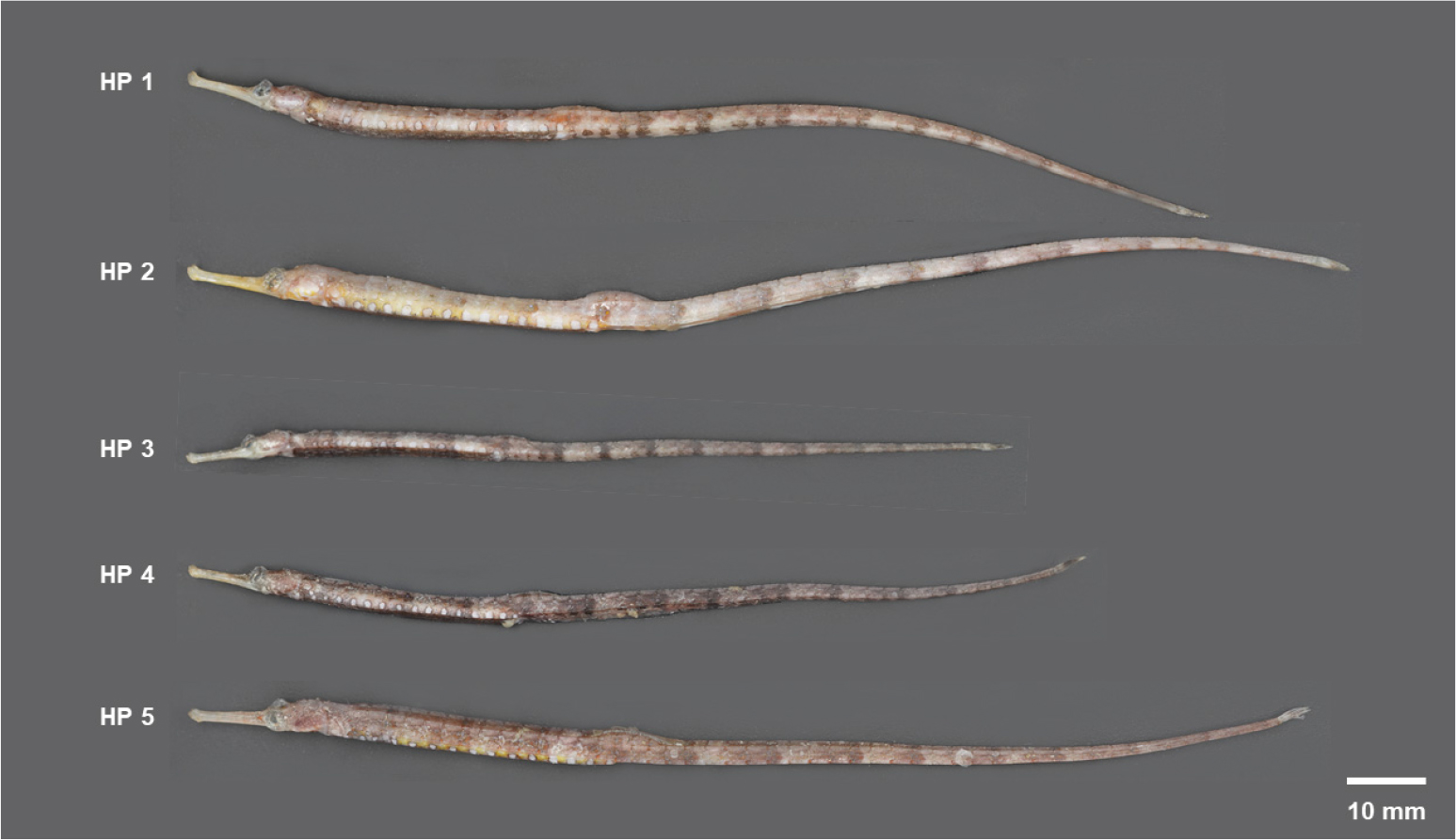
Five adult specimens (HP 1–5) were collected on August 17, 2025, off the eastern waters off Jeju Island (33°30′ N, 127°44′ E; depth 110 m). Sampling was conducted using a benthic dredge (opening 115×86 cm, net length 266 cm, mesh size 1 cm) aboard the R/V Onnuri. Upon collection, the specimens were immediately frozen at –20°C. In the laboratory, the specimens were thawed, and their total lengths and external morphology were photographed. Meristic characters were recorded by counting trunk rings (pectoral fin base to anus) and tail rings (anus to caudal fin base).
2.2 Molecular Analysis
Genomic DNA (gDNA) was extracted from muscle tissue using the AccuPrep® Genomic DNA Extraction Kit (Bioneer, Korea). For taxonomic identification, a partial region of the mitochondrial cytochrome c oxidase subunit I (COI) gene was amplified using the primers FishF2_t1 and FishR2_t1 (Ward et al. 2005). PCR amplification consisted of an initial denaturation at 94°C for 3 min, followed by 35 cycles of denaturation at 94°C for 30 s, annealing at 52°C for 40 s, and extension at 72°C for 1 min, with a final extension at 72°C for 7 min. The PCR products were purified using the MagListoTM 5M PCR Purification Kit (Bioneer, Korea) and sequenced on an ABI 3730xl DNA Analyzer (Thermo Fisher Scientific, USA). The newly generated COI sequences were deposited in NCBI GenBank (Accession numbers: HP 1, PX658561; HP 2, PX658565; HP 3, PX658563; HP 4, PX658562; HP 5, PX658564).
2.3 Phylogenetic and Biogeographical Analysis
Reference COI sequences for the family Syngnathidae were obtained from NCBI GenBank (Clark et al. 2016). Sequences were aligned using CLUSTAL W (Thompson et al. 1994), and both intraspecific and interspecific Kimura 2-parameter (K2P) distances were calculated using MEGA-X (Kumar et al. 2018). Phylogenetic relationships were reconstructed using the Maximum Likelihood (ML) method based on the best-fit GTR+G+I model (Nei and Kumar 2000). To assess the biogeographical range of H. punctatus, adult distribution data (FishBase 2025; Takata and Sasaki 2001) were compared with previous larval occurrence records (Kim et al. 2006).
3. Results and Discussion
The total lengths (TL) of the specimens examined in this study ranged from 103.2 to 147.3 mm (mean 127.3 mm). Morphologically, 14–15 ocelli were observed on the trunk, and 13 bands were arranged at regular intervals from the trunk to the tail. No cirri were present on the dorsal trunk, and the ring count was consistent across all specimens at 14+35. These meristic and morphological characters are consistent with the diagnostic characters previously reported for Halicampus punctatus (Kamohara 1952; Takata and Sasaki 2001; Kim et al. 2006). Based on these features, the specimens were identified as H. punctatus (Fig. 1).
The Kimura 2-parameter (K2P) distances of the partial COI sequences (614 bp) obtained from the five specimens were very low, ranging from 0.00% to 0.53% (mean 0.35%). This value is lower than the average intraspecific variation of 0.49% reported for the family Syngnathidae (Zhang et al. 2017), supporting all Korean individuals (HP 1–5) belong to the same species.
In the maximum-likelihood (ML) phylogenetic tree constructed using the present specimens and 32 reference COI sequences of Syngnathidae, all five Korean specimens formed a monophyly with Halicampus macrorhynchus (LC867271), supported by a bootstrap value of 89% (Fig. 2). The intraspecific K2P distance among the Korean specimens (HP 1–5) was very low, ranging from 0.00% to 0.53% (a difference of 0 to 3 bp), strongly supporting their identification as a single species. However, the K2P distance between H. macrorhynchus (LC867271) and Korean individuals (HP 1–5) was 14.4% (a difference of 82 bp); this exceeds the average 12.53% in syngnathids (Zhang et al. 2017) and makes a definitive congeneric conclusion based solely on this value difficult. Furthermore, another sequence registered as H. macrorhynchus (KY066101) showed a K2P distance ranging from 20.26% to 20.76% (a difference of 115 to 118 bp) from Korean specimens—approaching the average confamilial distance of 24.06% (Zhang et al. 2017)—and did not cluster with the H. macrorhynchus (LC867271). The K2P distances between Korean specimens HP 1–5 and other Halicampus species, such as H. brocki (KY066098), H. dunckeri (KY066099), and H. nitidus (KP194446), ranged from 22.75% to 23.47% (a difference of 129 to 134 bp), averaging 22.96%. Moreover, the interspecific distances among these reference species ranged from 17.72% to 23.53% (mean 19.91%), corresponding to a difference of 101 to 134 bp. These results indicate that while our specimens are phylogenetically closest to the H. macrorhynchus (LC867271), the genus Halicampus as a whole does not form a monophyletic group.
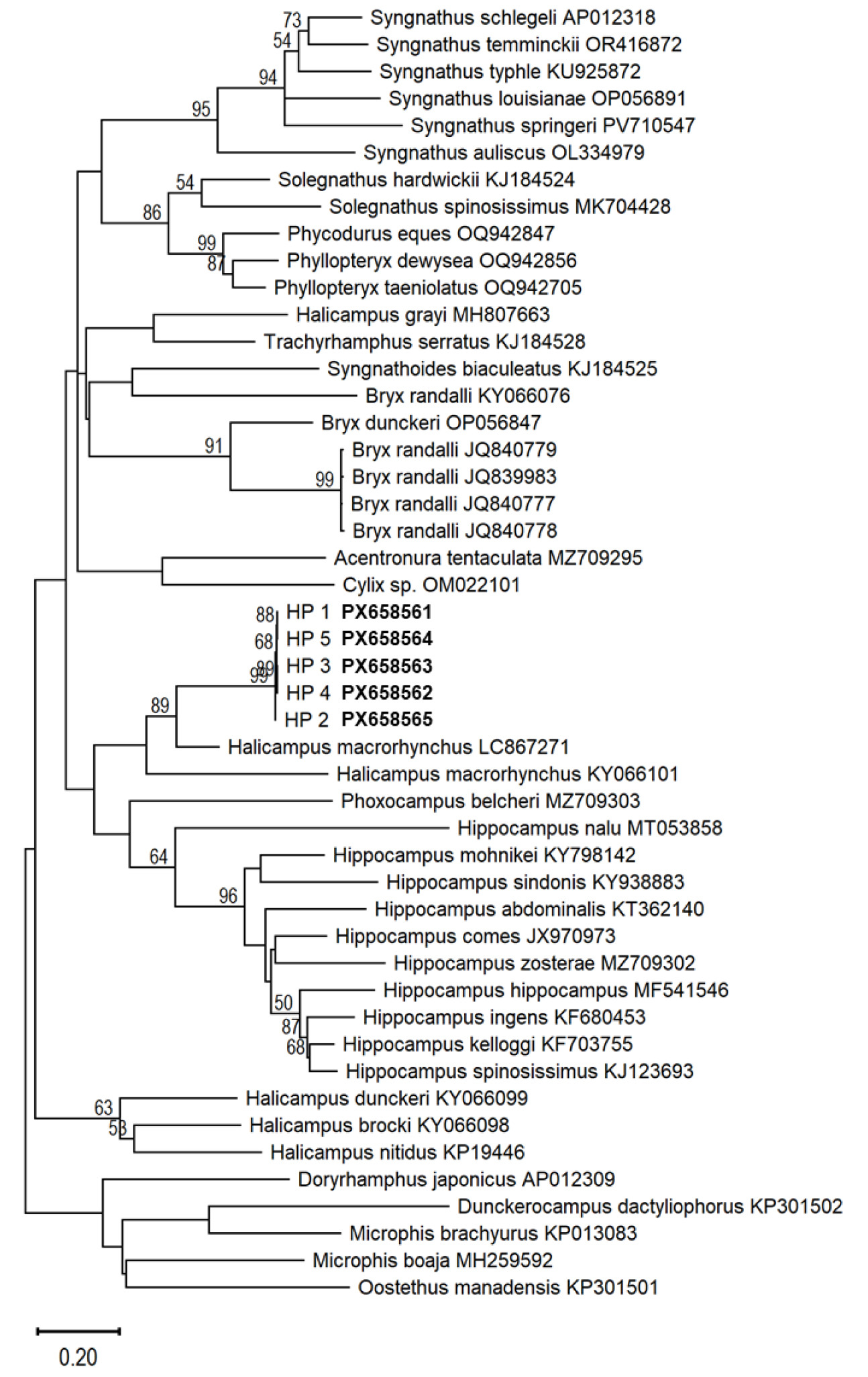
Fig. 2
Maximum Likelihood (ML) phylogenetic tree based on mitochondrial cytochrome c oxidase subunit I (COI) sequences from the five Halicampus punctatus specimens (HP 1–5; see Fig. 1) and 43 reference species of Syngnathidae. Numbers at nodes indicate bootstrap support values (>50%) calculated from 1,000 replicates. The scale bar indicates the number of nucleotide substitutions per site.
Notably, the K2P distance between the two sequences registered as H. macrorhynchus (LC867271 and KY066101) was 18.61%, significantly exceeding the average congeneric distance of 12.53% (Zhang et al. 2017). This discrepancy suggests that these two sequences may be from different species warranting a taxonomic revision of H. macrorhynchus. Furthermore, these findings imply that the current publicly available COI sequences for Halicampus are insufficient to clearly define the genetic boundaries of the genus.
Although H. punctatus has been considered a species with a distribution restricted to the waters around Japan (Dawson 1985; Senou 2002; Sonoyama et al. 2020), this study provides the first record of adult specimens from a depth of 110 m off the eastern waters off Jeju Island, Korea (Fig. 3). Previously, Kim et al. (2006) reported the occurrence of pelagic larvae of this species in the northwestern Korea Strait. Taken together, these records suggest that the distribution of H. punctatus is not limited to Japanese waters. Instead, this species is likely distributed across the broader region influenced by the Tsushima Warm Current and the Kuroshio Current.

Fig. 3
Map showing the distribution of Halicampus punctatus in the Northwest Pacific. The black triangle (▲) indicates the collection site of adult specimens (HP 1–5) in this study. Blue diamonds (◆) represent larval occurrences reported by Kim et al. (2006), and red circles (●) indicate previous adult records (Takata and Sasaki 2001; FishBase 2025).
In conclusion, this study presents the first record of adult H. punctatus in the waters around the Korean Peninsula. Simultaneously, our genetic analysis confirms that the genus Halicampus is non-monophyletic and highlights the ambiguity of species boundaries within the genus. To resolve the taxonomic system of this genus, further studies involving more extensive sampling and in-depth phylogenetic analyses based on mitogenome and nuclear gene markers are necessary.